Selectional Combinatorial GMDH
We will briefly describe a general idea of the Selectional Combinatorial GMDH algorithm according to [1], albeit using a somewhat different terminology, in context of multivariate smooth-function regression. Informally, the problem of regression by GMDH can be stated as follows: out of a set of measurements, model the dependent variable as a function of explanatory variables (regressors) optimally, according to some error criterion, where the function of dependency is represented by a GMDH network.
In order to define the manner in which GMDH networks get constructed, it is necessary to first introduce the basic building-blocks of GMDH networks and their connections. We discern two kinds of GMDH network primitives, see Fig. 1, the regressor nodes and the network nodes
, where
represents the total number of GMDH layers with the network nodes and
represents the beam search width i.e. the predetermined number of best partial solutions (network nodes) kept as candidates in layer
.
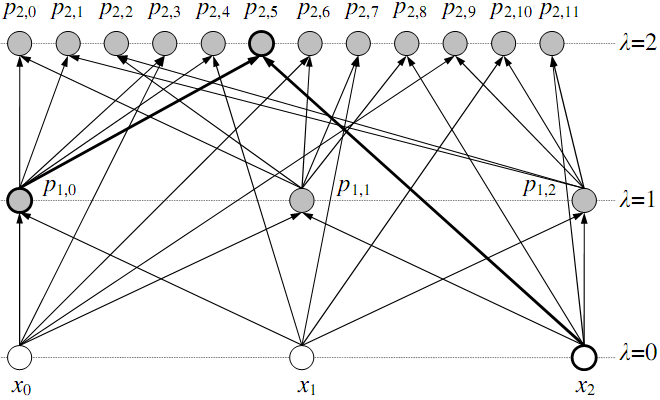 Figure 1. Building the GMDH networks
Figure 1. Building the GMDH networks
Generally, the network node is a two-input node with a second-order polynomial as the output nonlinearity. The regressor node
has no inputs, as it represents the regressor input into the network node. For example the network node
is constructed in the following way:
where and
can be either the regressor node or network node and
are the corresponding coefficients obtained by the polynomial regression.
The GMDH network is grown iteratively by inputting the nodes (either network or regressor nodes) to a new network node in a feedforward manner and performing low-order polynomial least-squares fitting to the dependent variable in order to obtain its coefficients. In that manner, the networks with complexities too small to capture the richness of dynamics of the process, which is modeled, can be used as inputs to a more complex network, better fitting the problem.
Still, finding the optimal structure of the network remains a problem due to the size of the space of possible solutions. To make the search feasible, a beam search is typically used in the GMDH algorithm. Keeping the algorithm tractable by such a constraint causes it to perform sub-optimally, but often a satisfactory sub-optimal solution suffices.
Let T and V be the training and the validation set, respectively, written in matrix form, with measurement instances concatenated as rows
where is the i-th regressor with samples in rows,
the dependent variable with samples in rows, with superscripts t and v denoting the test and the validation set, respectively, K is the number of regressors, M and N are the sizes of the training and validation sets, respectively.